Neurogenesis during whole-body regeneration
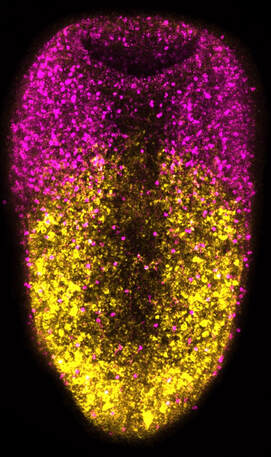
The complexity of the animal nervous system begs the question of how varied neural cell populations are generated. Further, understanding the mechanisms that effectively replace these neural populations in diverse animals provides a comparative framework to understand the evolution of neurogenesis. Most knowledge regarding the patterning and specification of neural populations comes from work focusing on embryonic development; however, limited work has been performed to identify mechanisms underlying the regeneration of entire nervous systems within adult animals. Vertebrates are capable of replacing select neural cell types using restricted progenitors; however, none are capable of whole brain regeneration. Several invertebrates are capable of whole-body regeneration, i.e., the capacity to replace any missing cell type, including neural cell types. Xenacoelmorpha is a clade thought to be a sister group to all other bilaterians and includes a number of highly regenerative animals. Of these animals, Hofstenia miamia, has been established as a model for mechanistic studies of whole-body regeneration. During my doctoral work, I identified a wound-induced differentiation trajectory for the recapitulation of a functional nervous system in H. miamia. To this end, I described the architecture and regeneration of the H. miamia nervous system, identified the heterogeneity of adult pluripotent stem cells and their putative trajectories including neural progenitors, and determined a gene regulatory network governing the early steps of neurogenesis during whole-body regeneration. Altogether, this work explored mechanisms governing neural specification and differentiation in H.miamia, allowing us to make inferences about how adult neurogenesis has evolved among animals.
Choanoflagellate multicellularity and interkingdom signaling
|
As a Research Technician in Nicole King’s lab at the University of California, Berkeley, I was drawn in because of this huge, unanswered question: How did animals become multicellular? This seemingly simple question was getting at the origins of animal multicellularity by reconstructing the basic genetic toolkit, understanding the ancestral functions of genes required for development, and teasing apart interspecies interactions. The model used in lab, S. rosetta, has an intricate life history with transitions between unicellular and multicellular stages and based on phylogenomics is the closest living relative of animals. I had the opportunity to tackle two different projects that both revolve around understanding the transition to multicelullarity in choanoflagellates and clarifying interspecies interactions. The first project centered around optimizing conditions for performing complementation and mapping crosses in S. rosetta to identify genes involved in the transition to multicellularity. The second project I worked on explored interspecies interactions between choanoflagellates and bacteria. A previous graduate student in the lab discovered that Vibrio fischeri induces sex in S. rosetta and I worked on various aspects of this interaction. I tested over 15 different Vibrio species to determine their ability to induce sex. I quantified the timeline during cellular and nuclear fusion using immunofluorescence and performed qPCR to determine meiotic gene expression. Both of these projects peaked my interest in understanding major transitions in animal complexity and how important it is to use a well placed, phylogenetically informative system to inform ancestral functions of genes and microbial interactions.
|
Nudibranch biogeography and color pattern evolution
|
My master’s thesis addressed the biodiversity, biogeography, and molecular systematics of tritoniid nudibranchs. Using molecular phylogenetics I explored the evolution of color and biogeography of tritoniid nudibranchs. I found that three well-supported clades within the genus Marionia are distinguished by their coloration including: translucence, blue spots, and longitudinal stripes. I performed an ancestral area reconstruction and determined that the family has biogeographical origins in the Indo-Pacific. This project reiterated the importance of biodiversity hotspots like the Indo-Pacific and emphasized the necessity of natural history collections.
|


